How a Jellyfish Changed the World
Glow worms in New Zealand masquerading as the night sky.
Being able to isolate a protein from nature is an important step in understanding how it functions. Sometimes, we are unable to observe how a protein works unless it is in an environment similar to where it is found in nature. For example, aequorin needs calcium to emit light, as this triggers a reaction that causes a light particle, called a photon, to be released. This property of aequorin was commonly used to study calcium concentrations inside our cells because calcium triggers numerous physiological responses including muscle contraction, blood clotting, and maintaining your heart’s rhythmic heartbeat. What makes GFP so unique is that it can emit light naturally on its own, as long as there is oxygen around.
In 1994, Nobel laureate Dr. Martin Chalfie1 demonstrated that it was possible for GFP to glow in both E. coli and in the worm Caenorhabditis elegans (C. elegans). Both these organisms are translucent, just like a jellyfish, so it made observing the glowing properties of this molecule relatively simple. The tricky part was inserting the DNA code of the GFP protein from the jellyfish into that of the both E. coli and C. elegans and hoping that this protein would form properly and then glow in a completely different organism. Chalfie’s lab had the ideal combination of tenacity and luck when performing their experiments, as is the case with many incredible scientific discoveries. But what is the purpose of being able to make bacteria or animals glow green? The exciting part about genetic editing with GFP is that you can put it beside any gene to act as a glowing flag, showing you where your gene of interest is building proteins in the organism. For Dr. Chalfie, he was interested in finding neurons in the brain that are responsible for sensing touch in his C. elegans worm model. This visually stunning and break-though work made the front cover of the world’s most highly-impactful science publication, Science! The use of GFP as a molecular marker exploded after this publication, evidenced by the 1500 sets of its genetic code (called a vector) that were sent out by the lab prior to its commercialization.
However, this is merely the beginning of GFPs scientific journey. It was Nobel Laureate, Dr. Roger Tsien1 that tapped into its true potential by not being satisfied with just one glowing protein. Although a green glowing protein is really cool, just imagine a glowing rainbow! You could tag many different genes at the same time to see how they work together, and coincidentally create both informative and pretty pictures. Dr. Tsien was interested to see if slightly mutating the protein could create different colours. These mutations could not be random because they could disrupt the proteins’ ability to glow. Being able to accurately manipulate GFP involves understanding its structure, which involves using x-ray crystallography. To briefly explain, x-ray crystallography involves having enough quantity of a molecular-sized substance, in our case GFP, to form a crystal where x-rays can be beamed through it. The diffraction, or way the x-rays spread out, after being beamed through the crystal provides scientists with data that they use to determine the molecular structure of the individual molecule that the crystal is made of. It turns out that GFP looks like a cylinder with a helix structure in its center. The chemical group responsible for the glow’s specific colour (called a chromophore) is deep in the center of the protein, so Dr. Tsien’s group were able to figure out what could be altered to cause different colours to glow. His lab was able to create blue, cyan, and yellow fluorescence proteins in addition to the natural-occurring green one.
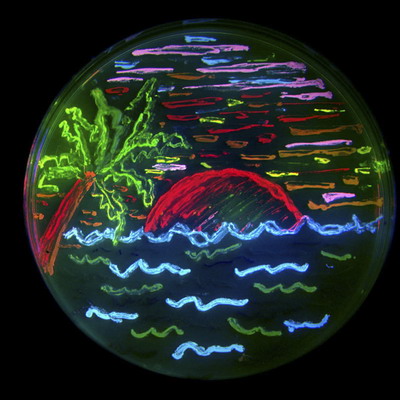
An example of a fluorescent protein rainbow.
Due to their similarity of their form, GFP and RFP can be used in similar experiments, thus effectively completing a fluorescent rainbow and providing scientists with an exciting labeling tool to work with. As of 2019, there are currently over 60 000 biomedical publications that cite using green fluorescent protein in their experiments, too many of which to be mentioned in this article, but here are a few awesome examples:
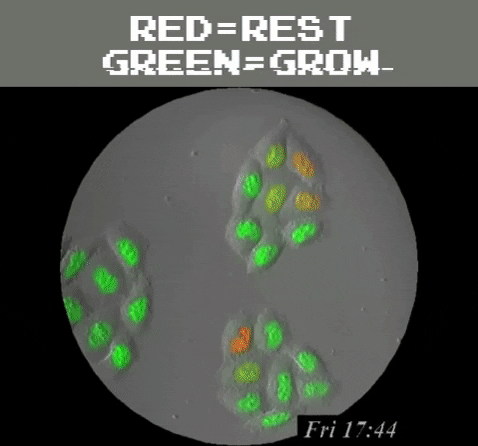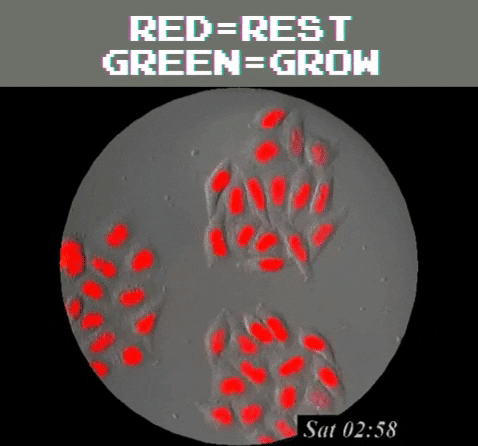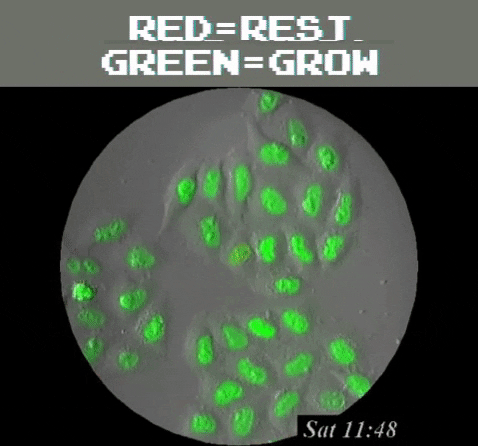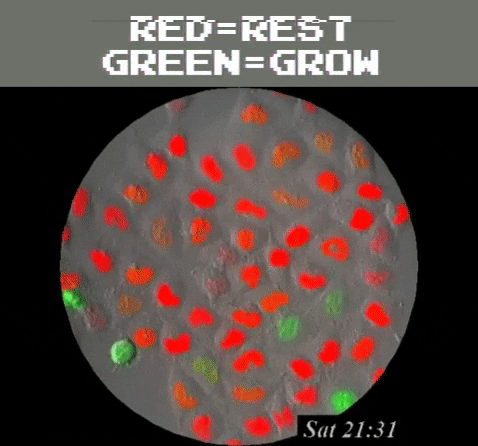
Amazingly, GFPs usefulness extend even beyond understanding the genetics and physiology of living creatures. It can be used as a tool to detect problematic metallic trace elements6 that contribute to environmental concerns. Of course, biology is still involved because a special strain of E. coli is used as a biosensor that emits green light once it detects the metallic trace elements. In general, GFP’s function as a label to determine the locations of specific proteins, track and monitor cell signals, figure out proteins’ lifespans, and see how molecules interact with each other, among many other uses, all harkens back to the observation of a humble, yet stunning, glowing jellyfish.
References/More Information
If you are interested in learning more about Nobel laureates that were awarded for working with GFP, check out the Nobel Prize in Chemistry 2008.
Matz MV, et al. Fluorescent proteins from nonbioluminescent Anthozoa species. Nature Biotechnology 1999 17: 969-973.
Chen P, et al. Predominant mode of human immunodeficiency virus transfer between T-cells is mediated by sustained Env-dependent neutralization-resistant virological synapses. Journal of Virology 2007 81: 12582-12595.
Kuchibhotla KV, et al. A beta plaques lead to aberrant regulation of calcium homeostasis in vivo resulting in structural and functional disruption of neuronal networks. Neuron 2008 59: 214-224.
Sakaue-Sawano A, et al. Visualizing spatiotemporal dynamics of multicellular cell-cycle progression. Cell 2008 132: 487-498.
Hurdebise Q, et al. Determination of zinc, cadmium, and lead bioavailability in contaminated soils at the single-cell level by a combination of whole-cell biosensors and flow cytometry. Sensors 2015, 15: 8981-8999.